
Lawlor Lab
Unraveling the mysteries of Ewing sarcoma
The goal of the Lawlor Lab is to discover how hijacking of normal developmental biologic programs contributes to the origin and progression of childhood cancer. Our research is focused on Ewing sarcoma, an aggressive bone and soft tissue tumor for which new cures are needed. Through investigations of tumor biology, we are identifying new therapeutic opportunities. Current areas of interest are tumor heterogeneity, epigenetic and metabolic reprogramming, and discovering how these processes contribute to tumor metastasis.
Featured Research
Tumor heterogeneity
Sarcomas are complex tissues that are comprised of both cancer cells and normal cells, as well as extracellular matrix. Crosstalk and collaboration among all components of the tumor is necessary for local tumor growth as well as metastasis to distant sites. Importantly, our research has shown that Ewing sarcoma tumor cells are very dynamic and they change phenotype rapidly in response to signals from their microenvironment. It is clear from our studies, as well as others, that these changes in cancer cell phenotype contribute to the tumor heterogeneity that drives disease relapse, metastasis, and emergence of drug resistant disease. Using diverse tumor model systems and state-of-the-art genomic, epigenomic, and functional assays, we are elucidating the molecular mechanisms that cause tumor cells to change states.
Our most recent work has focused on tumor microenvironment crosstalk and how Wnt/beta-catenin and TGF-beta signaling in discrete tumor-cell subpopulations collaborate to alter the local extracellular matrix, promoting tumor angiogenesis and disease progression.
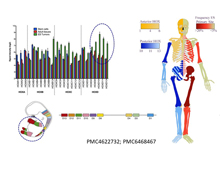
Enhancer reprogramming of posterior HOXD genes is an epigenetic hallmark of Ewing sarcoma. We have shown that hijacking of HOXD13 regulation specifically coopts developmental transcription programs that are essential for tumorigenicity.
Epigenetics
Ewing sarcomas are characterized by the presence of gene fusions, most commonly EWS-FLI1, often without evidence of any other gene mutations. As such, changes in cell phenotype that promote metastatic progression are largely driven by non-genetic mechanisms.
We are studying how disruption of normal epigenetic regulation of developmental programs leads to tumor initiation and tumor progression. In particular, we are focused on how epigenetic hijacking of developmental transcription factors such as HOX genes leads to malignant transformation of normal stem cells. In addition, we are studying how these processes work in established sarcoma cells to switch chromatin states allowing cells to adapt to new environments and to stress.
Understanding the fundamental biology of epigenetic processes in Ewing sarcoma will lead to new treatment strategies that employ less toxic, tumor-targeted drugs (i.e. inhibitors of bromodomain proteins, HDACs, menin, and other epigenetic modifying agents).
Metabolic reprogramming
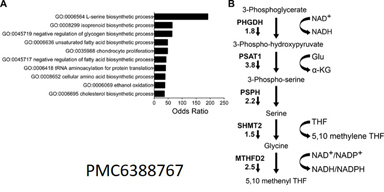 Inhibition of the scaffolding protein menin in Ewing sarcoma leads to metabolic rewiring, with specific loss of serine synthesis pathway (SSP) gene expression and activity. This creates a metabolic vulnerability that we hypothesize can be therapeutically exploited.
Inhibition of the scaffolding protein menin in Ewing sarcoma leads to metabolic rewiring, with specific loss of serine synthesis pathway (SSP) gene expression and activity. This creates a metabolic vulnerability that we hypothesize can be therapeutically exploited.The process of rapid proliferation and tumor expansion exposes tumor cells to multiple sources of internal and external stress. Tumor cells have adapted ways to maintain cellular homeostasis that allow them to survive and thrive under conditions that would destroy less resilient cells. Among the most important of these adaptation mechanisms is metabolic reprogramming. By rewiring their metabolic pathways, cancer cells are able to maintain redox and amino acid homeostasis and epigenetic stability, while generating the extra energy and biomaterials that are necessary to sustain their growth.
Our current studies of Ewing sarcoma are investigating how serine biosynthesis and amino acid flux contribute to tumor progression. Importantly, these studies have identified critical connections between the epigenome and metabolic reprogramming that we hypothesize are necessary for metastasis.
Partnership Opportunities

Elizabeth Lawlor, MD, PhD
Dr. Elizabeth Lawlor completed her MD at McMaster University and her clinical training in pediatric hematology-oncology and bone marrow transplantation at BC Children’s Hospital in Vancouver, Canada. Her passion to discover new cures for childhood cancer led her back to school, where she completed her PhD in pathology and laboratory medicine at the University of British Columbia. She then moved to the University of California, San Francisco to finish her post-doctoral training and then began her career as an independent investigator in 2004 at Children’s Hospital Los Angeles and the University of Southern California. In 2010, Lawlor moved to the University of Michigan, where her lab continued to focus its research on basic and translational studies of pediatric solid tumors, in particular Ewing sarcoma. While at Michigan, she also dedicated her energies to a second passion – graduate and post-graduate education – serving as director of the Cancer Biology PhD graduate and T32 training programs and as associate director of education and training at the Rogel Cancer Center.
Lawlor joined Seattle Children’s and the Ben Towne Center for Childhood Cancer and Blood Disorders Research in June 2020 and currently serves as associate director for the center. She is also a professor of Pediatrics at the University of Washington and adjunct professor in the Department of Laboratory Medicine and Pathology. The overall goal of her research is to discover and define differences between normal developmental and sarcoma biology that will enable discovery of targeted therapeutics that are less toxic to growing children.
Lawlor’s lab has been continuously funded by grants from the National Institutes of Health/National Cancer Institute, American Association for Cancer Research/Stand Up to Cancer, Alex’s Lemonade Stand Foundation, St. Baldrick’s Foundation, the V Foundation, as well as numerous smaller funding agencies. As a dual-trained MD and PhD physician-scientist, she is committed to educating the next generation of cancer researchers and to creating a diverse pipeline of physicians and scientists who will lead us to a better tomorrow for children who are diagnosed with cancer.
-

Katherine Braun, PhD
Research Scientist
Katherine Braun received her BS in biochemistry from the University of California, Davis, and her PhD in biochemistry from the University of Iowa. Her work as a graduate student and a postdoctoral scholar at Cold Spring Harbor Laboratory focused on understanding the mechanism of DNA replication in eukaryotes. In subsequent postdoctoral work at Fred Hutchinson Cancer Research Center, she studied cell cycle regulation in yeast. As a research scientist at the University of Washington, she studied the regulation of gene expression in response to changing nutrient conditions in yeast. She moved to the Fred Hutchinson Cancer Research Center to study chemoresistance in breast cancer using patient-derived organoids as a senior research scientist. She joined Seattle Children’s Research Institute in 2020 as a senior research scientist and her research focuses on understanding the role of Menin in modulating cell state and metastasis in Ewing sarcoma.
-

Eric Bueter, PhD
Research Scientist
Eric completed his B.S. in Chemistry at Wake Forest University before earning a Ph.D. in Cancer Biology from the University of Chicago. His graduate research, under the mentorship of Dr. Russell Szmulewitz, focused on characterizing a novel mechanism of resistance to anti-androgen therapy in prostate cancer. He subsequently pursued a postdoctoral fellowship in the SEPS lab at Seattle Children’s where he utilized an emerging proteomic technology, quantitative multiplex immunoprecipitation, to interrogate chimeric antigen receptor (CAR) signal transduction at network scale. Following a brief stint in industry, Eric returned to Seattle Children’s in 2025 to join the Lawlor lab as a senior scientist. He is currently investigating the molecular mechanisms of metastasis in Ewing sarcoma with the goal of finding unique vulnerabilities that can be exploited for therapeutic benefit.
-

Postdoctoral Researcher
Shireen received her MD from the University of Toledo College of Medicine in 2016. She completed her pediatric residency at UPMC Children’s Hospital of Pittsburgh in 2019, and her pediatric hematology/oncology fellowship at the University of Washington in 2022. She is currently completing her postdoctoral research fellowship. Her particular research focus is on preclinical testing and elucidating resistance mechanisms of the bromodomain inhibitor, BMS-986158, in Ewing sarcoma. She is additionally interrogating how anthracyclines, a class of drugs used widely in Ewing sarcoma and many pediatric cancers, disrupt key epigenetic and transcriptional networks. While not in the lab, she is an attending physician in the Cancer and Blood Disorders Center at Seattle Children’s Hospital and treats pediatric, adolescent and young adult patients with solid tumors.
-

Nicolas Garcia, MS
Research Scientist
Nicolas was born and raised in France, received his degrees from the University of Nice Sophia Antipolis, and worked at the ENS Lyon before moving to the United States in 2010. He conducted research on Flavivirus at the Center for Vaccine Research of Pittsburgh and then moved to the West Coast at the Fred Hutchinson Cancer Research Center to work on T-cell therapy for solid and liquid tumors before joining Seattle Children’s.
-

Aakanksha Jha, PhD
Invent Scholar, Seattle Children’s Research Institute
Aakanksha got her BE from Gujarat Technological University, her MS from the University of Florida and her PhD from the University of Florida. She engineers pre-clinical hydrogel models for Ewing sarcoma. Specifically, she is modeling the TME for the hydrogels to serve as a drug screening platform and for identifying other biological markers.
-

Jonah Valenti
Research Scientist
Jonah earned his BS in general biology and oceanography from the University of Washington in 2024. His work is on processing data generated by experiments within the lab, aiming to establish pipelines to facilitate analysis for future studies.
-

Stephanie Walter
Research Technician
Stephanie earned her BS in cell and molecular biology from Seattle University in 2023. She works on projects focused on understanding Menin’s role in metastasis in Ewing sarcoma, and the transcriptional and epigenetic effects of anthracyclines on Ewing sarcoma cells.
-

Emma Wrenn, PhD
Postdoctoral Researcher
Emma earned her BS from UC Santa Barbara and her PhD from the Molecular and Cellular Biology Graduate Program at UW/Fred Hutchinson in 2021. Her work focuses on how specific subgroups of Ewing sarcoma cells remodel the extracellular matrix, promoting disease progression and treatment resistance.
