Research Links Low Early Warning Signals in the Airway to Severe Viral-Triggered Asthma Attacks
Published
Featured Researchers
Summary
Researchers at the Center for Respiratory Biology and Therapeutics at Seattle Children’s Research Institute recently published findings in JCI Insight that reveal why certain children are more susceptible to life-threatening asthma attacks.
Asthma affects nearly 1 in 15 children in the U.S., but for some, the condition is far more than a chronic nuisance—it is life-threatening. While viruses, environmental factors and allergies are known triggers, the Debley Lab at the Center for Respiratory Biology and Therapeutics is uncovering the cellular "why" behind severe attacks. By examining bronchial epithelial cells, researchers have identified that certain children are uniquely susceptible to respiratory viruses like rhinovirus. These findings pave the way for a new era of personalized, proactive care for the most at-risk pediatric patients.
Jason S. Debley, MD, MPH, director, Center for Respiratory Biology and Therapeutics, shared insights from this latest research:
What makes this research on pediatric asthma and viral infections unique?
For children susceptible to severe asthma attacks, respiratory viruses such as the common cold (rhinovirus) or the flu often serve as the trigger for a cascade of events leading to emergency care and hospitalization. The focus of asthma management has traditionally been reactive, controlling inflammation and muscle tightness around the airways once an exacerbation begins.
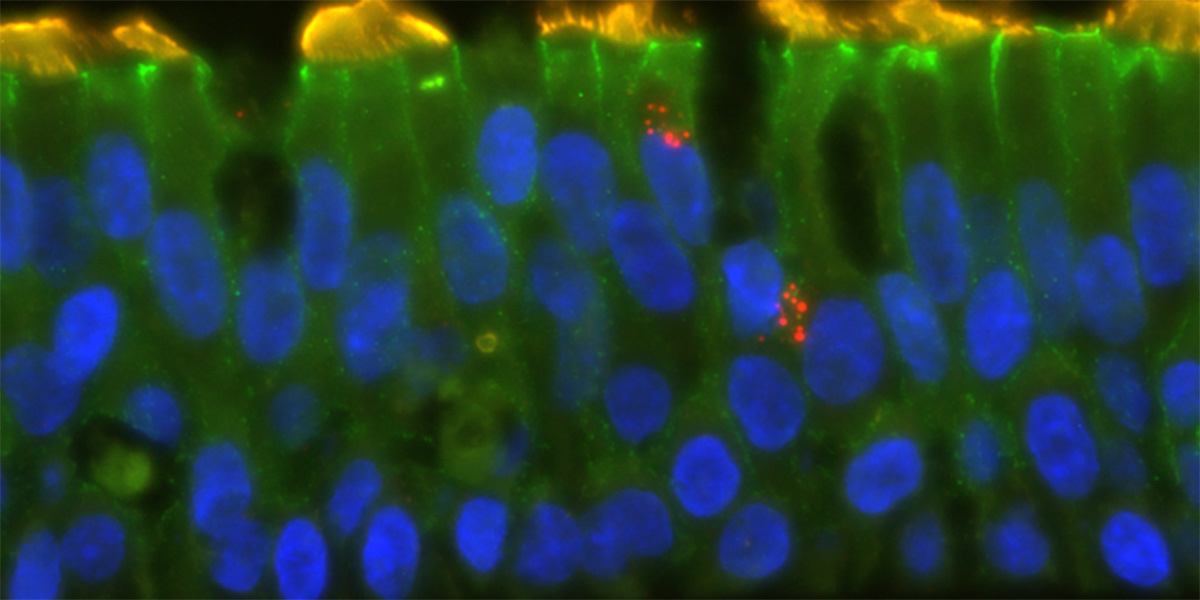
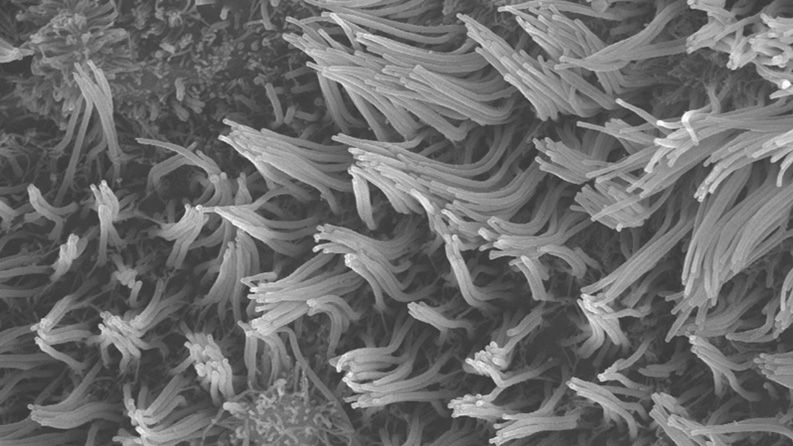 In one of the first studies of its kind, we used bronchial airway cells from children with and without asthma to demonstrate a direct mechanistic link between a child's baseline airway antiviral immune signaling and how their airway cells respond to viral infection. We found that when a child’s bronchial airway cells have lower baseline antiviral immune activity (interferon signaling), cold viruses replicate at a larger scale, leading to exaggerated inflammatory responses in children who are prone to severe asthma attacks.
In one of the first studies of its kind, we used bronchial airway cells from children with and without asthma to demonstrate a direct mechanistic link between a child's baseline airway antiviral immune signaling and how their airway cells respond to viral infection. We found that when a child’s bronchial airway cells have lower baseline antiviral immune activity (interferon signaling), cold viruses replicate at a larger scale, leading to exaggerated inflammatory responses in children who are prone to severe asthma attacks.
How could this research influence future asthma treatments and prevention?
This research opens two significant avenues for clinical and translational investigation. First is the potential for a predictive biomarker. If baseline airway interferon tone – the natural early antiviral signaling activity in airway cells that serves to defend against viral infection and warn neighboring cells –is a reliable indicator of risk, a straightforward method for measuring it in the airways could become a powerful tool to identify children with asthma at risk for severe attacks. This would allow clinicians to identify children who are most vulnerable to severe, virus-triggered exacerbations long before the high-risk respiratory virus season begins.
The second, and perhaps more transformative, implication is therapeutic. The data points toward a novel prophylactic strategy: boosting the airway's baseline antiviral defenses either preemptively or at the very first sign of infection. Interventions such as localized (e.g., inhaled), interferon-based treatments could boost antiviral signaling in the airway, transforming a vulnerable airway into a resilient one. This model of care would reposition our efforts to prevent the asthma attack from ever starting, rather than managing it after the fact.
The findings suggest that measuring baseline airway interferon tone to identify antiviral immune readiness in children's airways might help identify those at higher risk for severe asthma attacks triggered by viruses. Interferon-based treatments could potentially be used to enhance antiviral defenses early, offering a proactive approach rather than reactive treatment.
What are the future directions for this research?
Advancing this concept from the laboratory to clinical practice requires a coordinated and rigorous research effort. Our immediate priorities include:
- Developing accessible diagnostic tools. A central challenge is to create simple, non-invasive methods to measure baseline airway interferon tone in a clinical setting. This work is essential for translating the concept of risk stratification into a practical reality for pediatricians, pulmonologists, and allergists.
- Investigating the underlying biology. Key questions remain about the etiology of low airway interferon tone. We aim to understand why certain children have this reduced baseline antiviral immune activity, how this state changes over time from infancy through childhood and whether it can be safely and effectively modified.
- Validating and expanding our findings. It is critical to confirm these results in larger cohort and even more ethnically and socially diverse cohorts of children with asthma to ensure the applicability of this mechanism across different patient populations.
The long-term vision is to create a future where pediatric asthma care is deeply personalized and overwhelmingly preventative. By identifying at-risk individuals and employing targeted, immune-boosting therapies during high-risk seasons, we can aim to drastically reduce the burden of severe asthma attacks. This means fewer emergency visits, fewer hospitalizations and fewer missed days of school for children, aligning with our mission to help every child live the healthiest and most fulfilling life possible.
What collaborations and resources supported this research?
This research was made possible through funding by the National Institutes of Health and involved collaborations with the Benaroya Research Institute and the University of Washington.
Clinical characterization of children with asthma who participated in the study and all of the airway epithelial cell laboratory experiments were conducted by Dr. Jason Debley’s team in the Debley Lab at Seattle Children's Research Institute’s Center for Respiratory Biology and Therapeutics, with bioinformatics analyses supported by Basilin Benson and Drs. Naresh Doni Jayavelu and Matthew Altman at the Benaroya Research Institute Genomics Core.


