Seattle Children’s Research Institute’s Genomics and Spatial Biology Collaborative Laboratory provides access and specialized support for spatial biology research including transcriptomics, proteomics, genomics and related methodologies.
Genomics Equipment and Services
The PacBio Revio is a state-of-the-art long read sequencer that provides ultra high quality reads while also providing 5mC and 6mA calls by default on native (PCR-free) preps. PacBio’s Iso-Seq + Kinnex enables high yield full-length isoform sequencing at scale. Services offered by the CoLab are:
DNA library prep + sequencing:
- Whole genome
- Amplicon
Iso-Seq + Kinnex library prep + sequencing:
- RNA QC, cDNA synthesis, & Kinnex array formation
Sequencing only:
- Includes library QC + ABC preparation only
The CoLab provides accurate sizing for DNA and RNA of any length using the Agilent Femto Pulse, LightBench Discover and TapeStation 4200. We provide nanodrop and Qubit services for quantification.
The Illumina NextSeq 2000 is a desktop mid-throughput sequencer capable of producing between 100M and 1.8B reads per flow cell and read lengths up to 2x300bp.
The nCounter SPRINT Profiler provides an efficient, benchtop solution for digital gene expression, allowing for the direct, multiplexed counting of up to 800 targets without the need for cDNA synthesis or amplification. Its streamlined, all-in-one workflow is optimized for speed and flexibility, capable of processing 12 samples per run in approximately six hours even from challenging sources like FFPE tissue.
The Mission Bio Tapestri uses microfluidic droplet technology to deliver a high throughput single-cell genomics workflow for targeted DNA sequencing. The Tapestri integrates SNV, CNV, and protein data from the same cell.
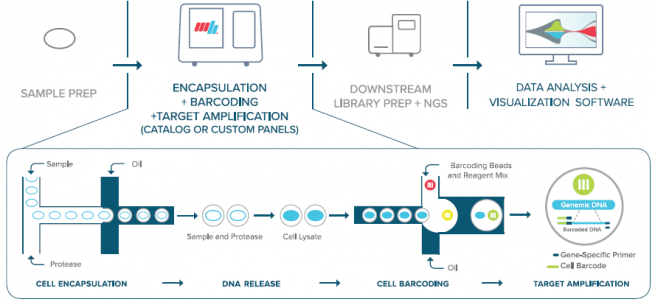
Image source: Mission Bio
Spatial Biology Equipment and Services
The Genomics and Spatial Biology CoLab facilitates the measurement of proteins and transcripts in tissue sections using the NanoString GeoMx Digital Spatial Profiler (DSP) platform. Digital spatial profiling preserves spatial information and allows transcriptomic and proteomic quantification within defined regions of fixed tissue, but it does not provide single-cell resolution. This technology is particularly suited for the characterization of rare cell types and specific microenvironments within tissue. For more info, see the original paper describing GeoMx DSP technology and a recent publication from Seattle Children's Research Institute researchers applying DSP to liver-stage malaria.
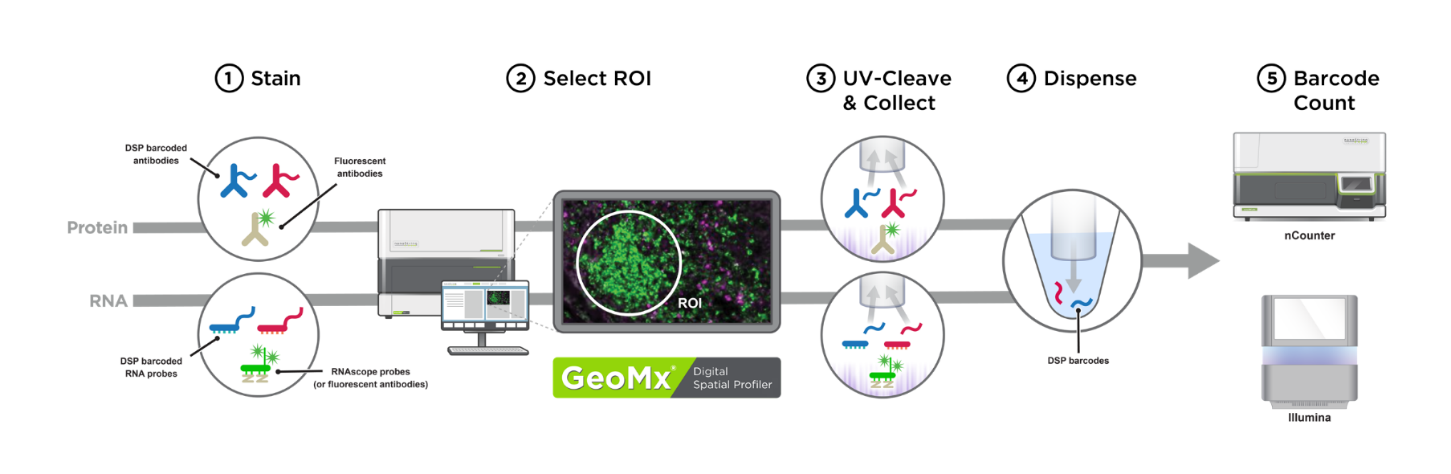
Image source: NeoGeomics Laboratories
The CosMx Spatial Molecular Imager (SMI) is a single-cell imaging platform that enables high-plex probe-based in situ analysis for spatial multiomics applications with formalin fixed paraffin embedded (FFPE) tissue samples at subcellular resolution. This instrument provides flexible and robust spatial single-cell imaging for applications of oncology, neuroscience, immunology, infectious disease and developmental biology research. CosMx has protein, RNA, and multiomic assays for human capable of profiling whole transcriptome and 1k universal or neuro panels for mouse tissue.
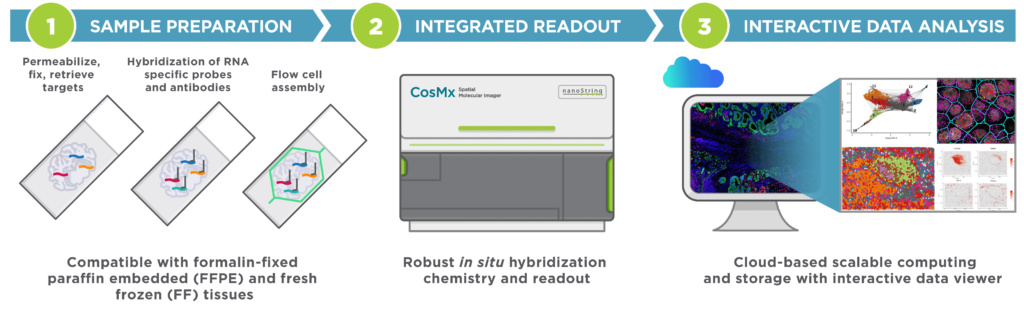
Image source: NanoString
The NanoString nCounter SPRINT Profiler combines the nCounter Pro Prep Station and Digital Analyzer into one instrument to allow for the digital examination of multiple pathways in a single tube. Output files include gene names and associated count numbers.

Image source: NanoString
Single Cell Services
The 10x Genomics Chromium platform uses advanced microfluidics to provide a versatile solution for profiling gene expression, immune repertoires, and cell surface proteins. Its diverse assay suite — including Single Cell ATAC, Multiome, and Fixed RNA Profiling — allows researchers to capture deep multiomic insights from fresh, frozen, or FFPE samples. By supporting both standard and high-throughput scales, it offers a standardized, reliable workflow for everything from pilot studies to massive cell atlases.
Parse Biosciences leverages combinatorial barcoding, significantly reducing the cost per cell while maintaining industry-leading data quality. Their unique upstream fixation workflow allows you to lock in biological states immediately at the point of collection, providing the flexibility to store and ship samples before processing them at scale. Parse provides kits for processing 10K, 100K, 1M, and 5M cells.
The Mission Bio Tapestri uses microfluidic droplet technology to deliver a high throughput single-cell genomics workflow for targeted DNA sequencing. The Tapestri integrates SNV, CNV, and protein data from the same cell. The CoLab provides services for DNA and DNA + protein workflows.
Request Services
Genomics and Spatial Biology CoLab services are available for internal and external users. To request services provided by the core, please visit the Genomics and Spatial Biology CoLab’s iLab page.
For information on how to access iLab, refer to this guide.
Genomics and Spatial Biology Rates
To view internal rates for genomics and spatial biology services, please visit this page.
Acknowledging Core Contributions
When citing the Genomics and Spatial Biology CoLab, we recommend the following language: Seattle Children's Research Institute Genomics and Spatial Biology Collaborative Laboratory, RRID:SCR_025490.
If you need more information for your grant, please contact the Genomics and Spatial Biology CoLab staff.
Access and Contact Information
Address for spatial biology and genomics services
Genomics and Spatial Biology CoLab
Seattle Children's Research Institute
1900 Ninth Ave., 7th Floor
Seattle, WA 98101
- General: [email protected]
- Rebecca Martin, PhD: [email protected]
- Mark Conner, BS: [email protected]
